High Throughput Glycan & Glycopeptide Identification Tool using Mass Spectrometry Data
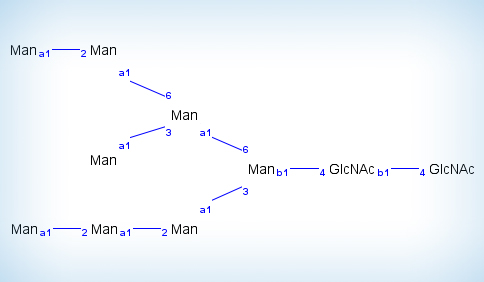
SimGlycan predicts structure of glycans from the MS/MS and multi-stage mass spectrometry (MSn) data. Furthermore, comprehensive support for resolving glycopeptides using LC-MS/MS glycopeptide data facilitates glycosylation studies. SimGlycan accepts experimental MS/MS data, matches them with its own database and finally generate a list of ranked candidate structures. Rank of each candidate indicates the proximity between the candidate structure and the experimental glycan. The rank is calculated based on a robust scoring mechanism that considers structure specific diagnostic ions observed in the experimental MS/MS spectra as well as the intensity of those observed peaks.
SimGlycan also supports multistage mass spectrometry (MSn) data analysis for isomers differentiation.
Workflows





- High Throughput Analysis of Native Glycan
- High Throughput Analysis of Reducing End Modified Released Glycans
- High Throughput Analyses of Permethylated Glycans
- Dual Reaction for Analytical Glycomics (DRAG) Analysis
- Glycan Quantitation using AminoxyTMT6 Reporter Ions
- High Throughput Glycopeptide Analysis