Oligo Design for Multiplex PCR & High Throughput SNP Genotyping and Analysis
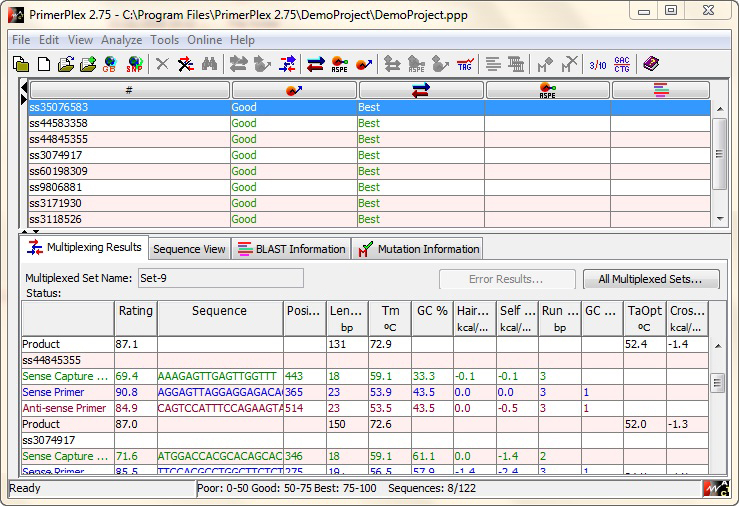
PrimerPlex is an efficient and sophisticated tool for designing oligos for multiplex assays. Multiplex assays facilitate amplification of multiple targets in a single reaction vessel, reducing both the time and cost of experimentation.
Primer design for multiplex PCR presents several challenges which include primer dimers, inability to separate amplicons with similar electrophoretic mobility and mis-priming due to nonspecific binding to non-target DNA templates. PrimerPlex uses proprietary algorithms to design optimal multiplex primer sets under uniform reaction conditions for over 100 sequences. The primer sets are identified after screening all the primers in a pool and minimizing Tm mismatches to ensure specific amplification and high signal strength. In this process, it analyzes millions of possible multiplex primer sets in a few seconds and presents a list of alternate sets. You can specify minimum product size differences amongst the set of designed primer pairs for better visualization of bands on a gel. To assure primer specificity, primers can be BLAST searched from PrimerPlex against any of the genomic databases available at NCBI.
Read More >>