References >> LAMP
LAMP - Loop Mediated Isothermal Amplification"LAMP" stands for Loop-mediated Isothermal Amplification. This technology was developed by Notomi et al. It is a very sensitive, easy and time efficient method. The LAMP reaction proceeds at a constant temperature using a strand displacement reaction.
Types of Primers used in LAMP
LAMP is characterized by the use of 4 different primers specifically designed to recognize 6 distinct regions of the target gene. The four primers used are as follows:
1. Forward Inner Primer (FIP): The FIP consists of a F2 region at the 3'end and a F1c region at the 5'end. The F2 region is complementary to the F2c region of the template sequence. The F1c region is identical to the F1c region of the template sequence.
2. Forward Outer Primer (FOP): The FOP (also called F3 Primer) consists of a F3 region which is complementary to the F3c region of the template sequence. This primer is shorter in length and lower in concentration than FIP.
3. Backward Inner Primer (BIP): The BIP consists of a B2 region at the 3'end and a B1c region at the 5'end. The B2 region is complementary to the B2c region of the template sequence. The B1c region is identical to the B1c region of the template sequence.
4. Backward Outer Primer (BOP): The BOP (also called B3 Primer) consists of a B3 region which is complementary to the B3c region of the template sequence.
Stages in Loop-mediated Isothermal Amplification
1. F2 region of FIP hybridizes to F2c region of the target DNA and initiates complementary strand synthesis.
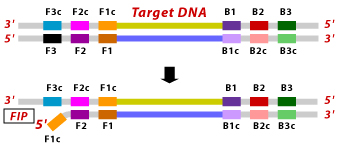
2. Outer primer F3 hybridizes to the F3c region of the target DNA and extends, displacing the FIP linked complementary strand. This displaced strand forms a loop at the 5' end.
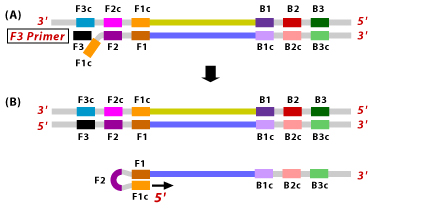
3. This single stranded DNA with a loop at the 5' end serves as a template for BIP. B2 hybridizes to B2c region of the template DNA. DNA synthesis is now initiated leading to the formation of a complementary strand and opening of the 5' end loop.
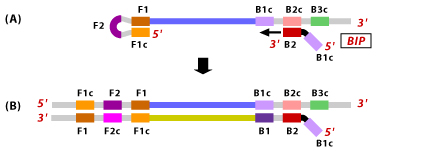
4. Now, the outer primer B3 hybridizes to B3c region of the target DNA and extends, displacing the BIP linked complementary strand. This results in the formation of a dumbbell shaped DNA.
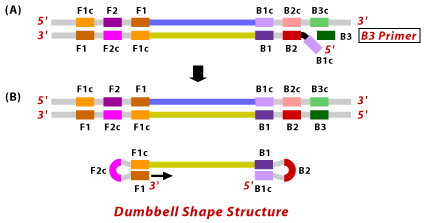
5. The nucleotides are added to the 3' end of F1 by DNA polymerase, which extends and opens up the loop at the 5' end. The dumbbell shaped DNA now gets converted to a stem loop structure. This structure serves as an initiator for LAMP cycling, which is the second stage of the LAMP reaction.

6. To initiate LAMP cycling, the FIP hybridizes to the loop of the stem-loop DNA structure. Strand synthesis is initiated here. As the FIP hybridizes to the loop, the F1 strand is displaced and forms a new loop at the 3' end.
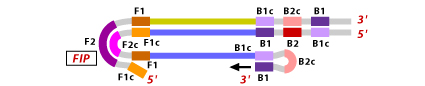
7. Now nucleotides are added to the 3' end of B1. The extension takes place displacing the FIP strand. This displaced strand again forms a dumbbell shaped DNA. Subsequent self-primed strand displacement DNA synthesis yields one complementary structure of the original stem loop DNA and one gap repaired stem loop DNA.
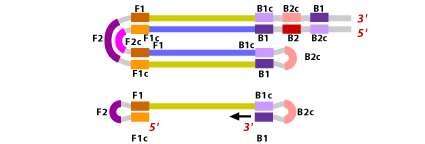
8. Both these products then serve as template for a BIP primed strand displacement reaction in the subsequent cycles. Thus, a LAMP target sequence is amplified 13 fold every half cycle.
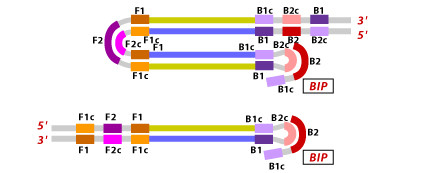
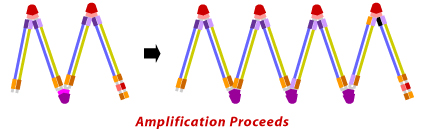
The final products obtained are a mixture of stem loop DNA with various stem lengths and various cauliflower like structures with multiple loops. The structures are formed by annealing between alternatively inverted repeats of the target sequence in the same strand.
Lamp Detection
In a LAMP assay, the reaction takes place in a single tube containing buffer, target DNA, DNA polymerase and primers. The tube is incubated at 64°C in a regular laboratory water bath or heat block that helps in maintaining a constant temperature. The amplified product can be detected by naked eye as a white precipitate or a yellow-green color solution after addition of SYBR green to the reaction tube.
Advantages
1. Amplification of DNA takes place at an isothermal condition (63 to 65°C) with greater efficiency.
2. Thermal denaturation of double stranded DNA is not required.
3. LAMP helps in specific amplification as it designs 4 primers to recognize 6 distinct regions on the target gene.
4. LAMP is cost effective as it does not require special reagents or sophisticated equipment.
5. This technology can be used for the amplification of RNA templates in presence of reverse transcriptase.
6. LAMP assay takes less time for amplification and detection.
Applications
1. LAMP is used in rapid diagnosis of viral, bacterial and parasitic diseases.
2. It helps in the identification of genus and species-specific parasites.
Efficient Primer Design for Loop-mediated Isothermal Amplification using LAMP Designer
LAMP Designer designs four primers along with two additional loop primers to identify six distinct regions. The software automatically interprets BLAST results and even avoids homologous regions on the sequence while designing primers to prevent non-specific amplification. The designed primers can also be BLAST searched to verify their specificity.
For details, please visit: http://www.premierbiosoft.com/isothermal/lamp.html



