Home >> Products >> Primer Premier
A Comprehensive PCR Primer Design Software
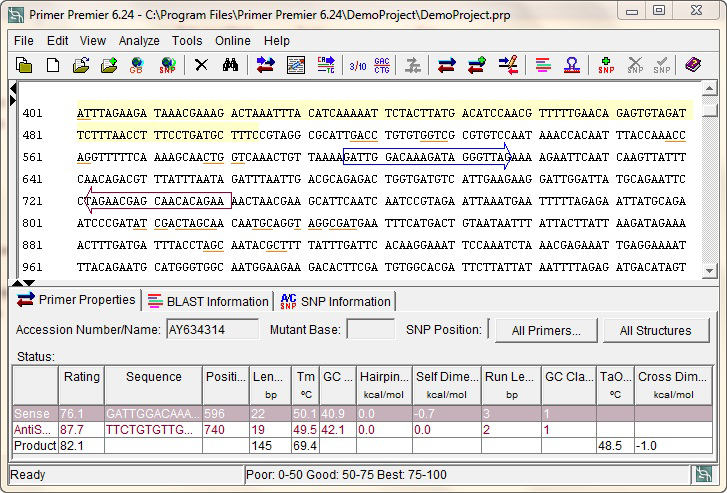
Primer Premier's search algorithm finds optimal PCR, multiplex and SNP genotyping primers with the most accurate melting temperature using the nearest neighbor algorithm. Primers are screened for secondary structures, dimers, hairpins, homologies and physical properties before reporting the best ones for your sequence, in a ranked order. Load the gene of interest from NCBI, select a search range, sit back and let Primer Premier pick the best possible primers for you.