Home >> Products >> MLPA® Designer
Designs Highly Specific MLPA® Assays
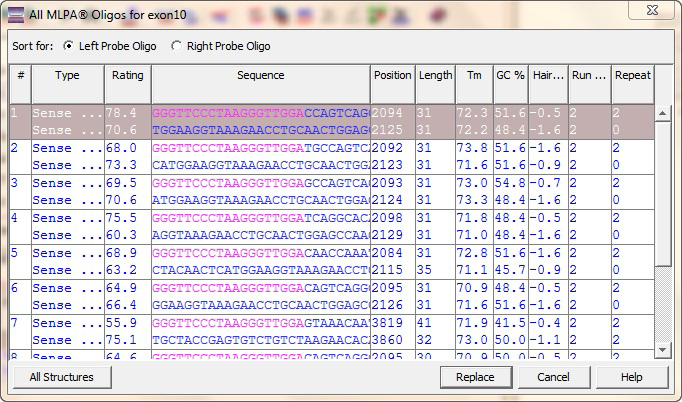
MLPA® Designer designs synthetic probes for MLPA (Multiplex Ligation dependent Probe Amplification) assays. The oligos are designed by avoiding regions of homologies making them highly specific. MLPA® Designer can be utilized to design oligos for both copy number detection and mutation studies. MLPA® is a simple, high throughput and easy to perform method developed by MRC-Holland that allows detection of DNA copy number changes of up to 50 sequences in a single reaction.